 Despite substantial progress in uncovering mechanisms of autoimmune disease pathogenesis and progression, the development of new, directed therapies for many autoimmune diseases has been marked by setbacks and controversies. For example, systemic lupus erythematosus (SLE), one of the top 10 causes of death for women between 15 and 45 years of age in the US, had >30 promising, strategically targeted biologics tested in clinical trials over the past several decades. Only one emerged as a new treatment for SLE. Other diseases, like neuromyelitis optica (NMO) and Sjögren’s syndrome, have no FDA-approved disease modifying drugs and woefully little research or clinical trials. These and other insights suggest that the path forward for autoimmune disease treatment must include a better understanding of molecular disease heterogeneity, immunologic subsets within and across diseases, the pharmacobiology of novel therapeutics, and the immunologic effects of background medications being used in trials with a polypharmacy design.
Despite substantial progress in uncovering mechanisms of autoimmune disease pathogenesis and progression, the development of new, directed therapies for many autoimmune diseases has been marked by setbacks and controversies. For example, systemic lupus erythematosus (SLE), one of the top 10 causes of death for women between 15 and 45 years of age in the US, had >30 promising, strategically targeted biologics tested in clinical trials over the past several decades. Only one emerged as a new treatment for SLE. Other diseases, like neuromyelitis optica (NMO) and Sjögren’s syndrome, have no FDA-approved disease modifying drugs and woefully little research or clinical trials. These and other insights suggest that the path forward for autoimmune disease treatment must include a better understanding of molecular disease heterogeneity, immunologic subsets within and across diseases, the pharmacobiology of novel therapeutics, and the immunologic effects of background medications being used in trials with a polypharmacy design.
Three projects in our group address this need. The Oklahoma Autoimmunity Center of Excellence (OACE; U19AI082714, UM1AI144292) focuses on understanding the pathogenic mechanisms of autoimmune disease flares within and across autoimmune disorders. As one of seven centers in the national ACE Network, the OACE specializes in planning and conducting clinical trials with integrated mechanistic studies. An experienced clinical and basic investigative team leads each OACE trial, and integrated mechanistic studies address hypotheses of disease pathogenesis, mechanisms of disease flare, biologic impact of the directed therapy, and predictors of treatment response.
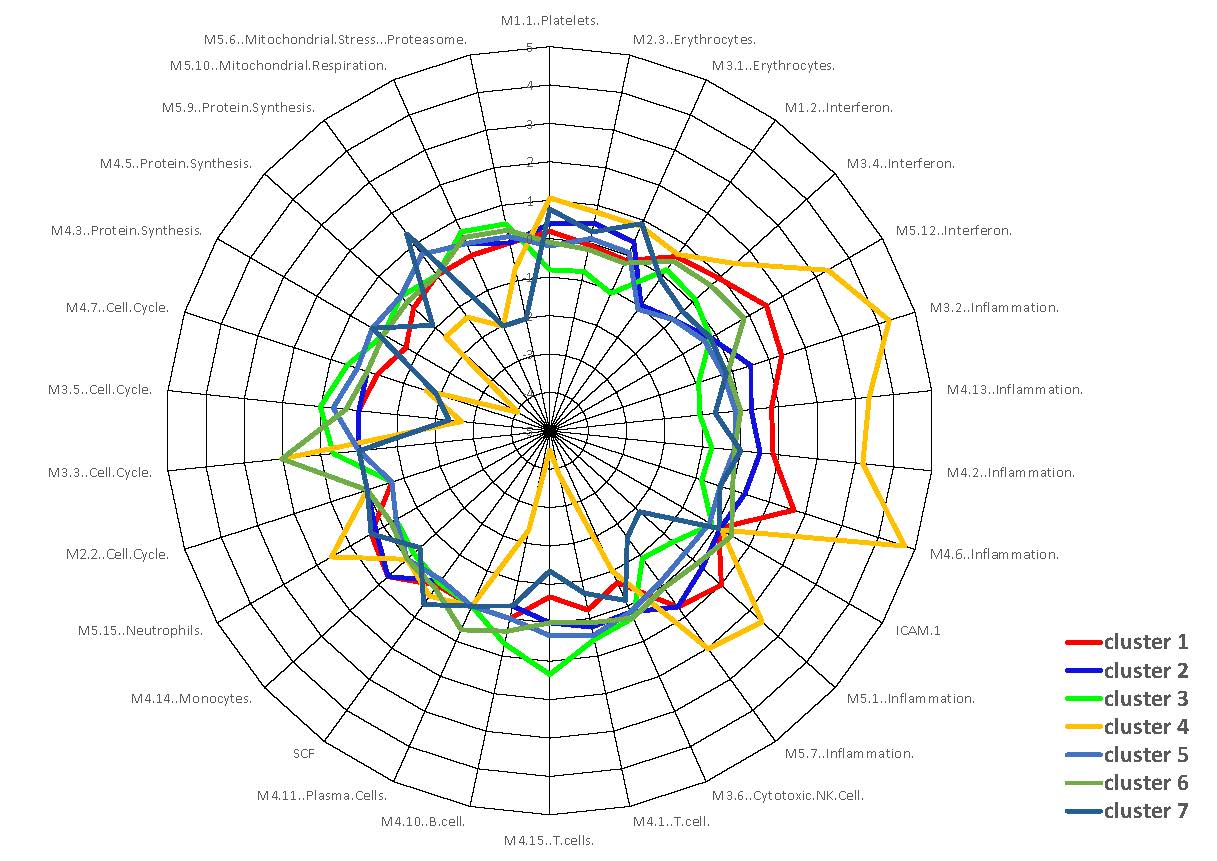
Our collaborative project through the ACE and our local research clinic deconstructs molecular heterogeneity and associated pathogenic mechanisms of disease in 7 subsets of SLE patients, which were identified through machine learning with high-dimensional data from the Oklahoma Lupus Cohort. This cohort follows 200 lupus patients with serial clinical assessments, gene expression profiling, soluble mediator assessment, and additional mechanistic analyses. Visits occur approximately every three months, thus providing a wealth of longitudinal information. Building on the identification of 7 molecular subsets, we are now testing hypotheses of specific molecular mechanisms through deep immunophenotyping and single cell technologies such as scRNAseq, CITE-seq, CyTOF and ChipCytometry.
The Oklahoma Rheumatic Disease Research Cores Center (ORDRCC; P30AR073750) focuses on enabling and applying precision medicine approaches to decipher the pathogenesis, prediction, and prevention of rheumatic diseases. As the foundation for this mission, the ORDRCC has developed some of the largest existing cross-sectional and longitudinal collections of SLE, Sjogren’s, and rheumatoid arthritis. We analyze these samples with high-dimensional cytometry, single-cell transcriptomics and proteomics, transcriptomics, and high-throughput testing of soluble biomarkers. We then use translational informatics approaches to visualize these complex datasets, understand disease mechanisms, and identify potential biomarkers that may inform future clinical trials, treatment selection, and disease monitoring. Through the RDRCC, we are also working on the development of new technologies for regular, at-home monitoring of immune function for rheumatic disease patients.
These studies are enabled by our outstanding clinical research infrastructure, which includes a CAP-certified biorepository, cutting-edge methodologies for mechanistic studies, unique cohorts/collections that represent diverse rural and minority communities, and a strong team of interdisciplinary investigators representing numerous clinical specialties and basic science disciplines.
Browse other areas:
Pre-clinical Autoimmunity
Mechanisms of Disease Flare
Improving Health Equity
Human Immunity to Infection and Vaccines